Understanding the decision tree structure¶
The decision tree structure can be analysed to gain further insight on the relation between the features and the target to predict. In this example, we show how to retrieve:
- the binary tree structure;
- the depth of each node and whether or not it’s a leaf;
- the nodes that were reached by a sample using the decision_path method;
- the leaf that was reached by a sample using the apply method;
- the rules that were used to predict a sample;
- the decision path shared by a group of samples.
import numpy as np from matplotlib import pyplot as plt from sklearn import tree from sklearn.datasets import load_iris from sklearn.model_selection import train_test_split from sklearn.tree import DecisionTreeClassifier
Train tree classifier¶
First, we fit a DecisionTreeClassifier using the load_iris dataset.
iris = load_iris() X = iris.data y = iris.target X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=0) clf = DecisionTreeClassifier(max_leaf_nodes=3, random_state=0) clf.fit(X_train, y_train)
DecisionTreeClassifier(max_leaf_nodes=3, random_state=0)
In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook.
On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
DecisionTreeClassifier(max_leaf_nodes=3, random_state=0)
Tree structure¶
The decision classifier has an attribute called tree_ which allows access to low level attributes such as node_count , the total number of nodes, and max_depth , the maximal depth of the tree. It also stores the entire binary tree structure, represented as a number of parallel arrays. The i-th element of each array holds information about the node i . Node 0 is the tree’s root. Some of the arrays only apply to either leaves or split nodes. In this case the values of the nodes of the other type is arbitrary. For example, the arrays feature and threshold only apply to split nodes. The values for leaf nodes in these arrays are therefore arbitrary.
Among these arrays, we have:
- children_left[i] : id of the left child of node i or -1 if leaf node
- children_right[i] : id of the right child of node i or -1 if leaf node
- feature[i] : feature used for splitting node i
- threshold[i] : threshold value at node i
- n_node_samples[i] : the number of training samples reaching node i
- impurity[i] : the impurity at node i
Using the arrays, we can traverse the tree structure to compute various properties. Below, we will compute the depth of each node and whether or not it is a leaf.
n_nodes = clf.tree_.node_count children_left = clf.tree_.children_left children_right = clf.tree_.children_right feature = clf.tree_.feature threshold = clf.tree_.threshold node_depth = np.zeros(shape=n_nodes, dtype=np.int64) is_leaves = np.zeros(shape=n_nodes, dtype=bool) stack = [(0, 0)] # start with the root node id (0) and its depth (0) while len(stack) > 0: # `pop` ensures each node is only visited once node_id, depth = stack.pop() node_depth[node_id] = depth # If the left and right child of a node is not the same we have a split # node is_split_node = children_left[node_id] != children_right[node_id] # If a split node, append left and right children and depth to `stack` # so we can loop through them if is_split_node: stack.append((children_left[node_id], depth + 1)) stack.append((children_right[node_id], depth + 1)) else: is_leaves[node_id] = True print( "The binary tree structure has nodes and has " "the following tree structure:\n".format(n=n_nodes) ) for i in range(n_nodes): if is_leaves[i]: print( " node= is a leaf node.".format( space=node_depth[i] * "\t", node=i ) ) else: print( " node= is a split node: " "go to node if X[:, ] " "else to node .".format( space=node_depth[i] * "\t", node=i, left=children_left[i], feature=feature[i], threshold=threshold[i], right=children_right[i], ) )
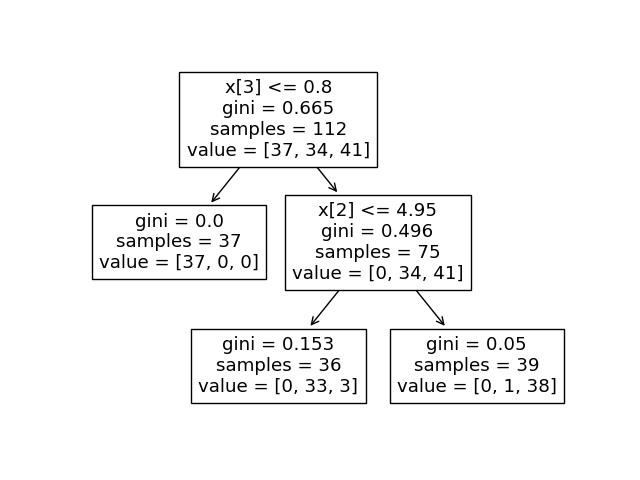
The binary tree structure has 5 nodes and has the following tree structure: node=0 is a split node: go to node 1 if X[:, 3]We can compare the above output to the plot of the decision tree.
Decision path¶
We can also retrieve the decision path of samples of interest. The decision_path method outputs an indicator matrix that allows us to retrieve the nodes the samples of interest traverse through. A non zero element in the indicator matrix at position (i, j) indicates that the sample i goes through the node j . Or, for one sample i , the positions of the non zero elements in row i of the indicator matrix designate the ids of the nodes that sample goes through.
The leaf ids reached by samples of interest can be obtained with the apply method. This returns an array of the node ids of the leaves reached by each sample of interest. Using the leaf ids and the decision_path we can obtain the splitting conditions that were used to predict a sample or a group of samples. First, let’s do it for one sample. Note that node_index is a sparse matrix.
node_indicator = clf.decision_path(X_test) leaf_id = clf.apply(X_test) sample_id = 0 # obtain ids of the nodes `sample_id` goes through, i.e., row `sample_id` node_index = node_indicator.indices[ node_indicator.indptr[sample_id] : node_indicator.indptr[sample_id + 1] ] print("Rules used to predict sample :\n".format(id=sample_id)) for node_id in node_index: # continue to the next node if it is a leaf node if leaf_id[sample_id] == node_id: continue # check if value of the split feature for sample 0 is below threshold if X_test[sample_id, feature[node_id]] threshold[node_id]: threshold_sign = " else: threshold_sign = ">" print( "decision node : (X_test[ , ] = ) " " )".format( node=node_id, sample=sample_id, feature=feature[node_id], value=X_test[sample_id, feature[node_id]], inequality=threshold_sign, threshold=threshold[node_id], ) )Rules used to predict sample 0: decision node 0 : (X_test[0, 3] = 2.4) > 0.800000011920929) decision node 2 : (X_test[0, 2] = 5.1) > 4.950000047683716)For a group of samples, we can determine the common nodes the samples go through.
sample_ids = [0, 1] # boolean array indicating the nodes both samples go through common_nodes = node_indicator.toarray()[sample_ids].sum(axis=0) == len(sample_ids) # obtain node ids using position in array common_node_id = np.arange(n_nodes)[common_nodes] print( "\nThe following samples share the node(s) in the tree.".format( samples=sample_ids, nodes=common_node_id ) ) print("This is % of all nodes.".format(prop=100 * len(common_node_id) / n_nodes))The following samples [0, 1] share the node(s) [0 2] in the tree. This is 40.0% of all nodes.Total running time of the script: ( 0 minutes 0.094 seconds)
Источник